Ch 12. 딥러닝 입문 (분류)
Part.3 Binary Classification
Load Dataset from sklearn
- In [1] :
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.preprocessing import StandardScaler
- In [2] :
from sklearn.datasets import load_breast_cancer
cancer = load_breast_cancer()
# 유방암 데이터 테이스
- In [3] :
df = pd.DataFrame(cancer.data, columns=cancer.feature_names)
df['class'] = cancer.target
df.tail()
- Out [3] :
| mean radius | mean texture | mean perimeter | mean area | mean smoothness | mean compactness | mean concavity | mean concave points | mean symmetry | mean fractal dimension | … | worst texture | worst perimeter | worst area | worst smoothness | worst compactness | worst concavity | worst concave points | worst symmetry | worst fractal dimension | class | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 564 | 21.56 | 22.39 | 142.00 | 1479.0 | 0.11100 | 0.11590 | 0.24390 | 0.13890 | 0.1726 | 0.05623 | … | 26.40 | 166.10 | 2027.0 | 0.14100 | 0.21130 | 0.4107 | 0.2216 | 0.2060 | 0.07115 | 0 |
| 565 | 20.13 | 28.25 | 131.20 | 1261.0 | 0.09780 | 0.10340 | 0.14400 | 0.09791 | 0.1752 | 0.05533 | … | 38.25 | 155.00 | 1731.0 | 0.11660 | 0.19220 | 0.3215 | 0.1628 | 0.2572 | 0.06637 | 0 |
| 566 | 16.60 | 28.08 | 108.30 | 858.1 | 0.08455 | 0.10230 | 0.09251 | 0.05302 | 0.1590 | 0.05648 | … | 34.12 | 126.70 | 1124.0 | 0.11390 | 0.30940 | 0.3403 | 0.1418 | 0.2218 | 0.07820 | 0 |
| 567 | 20.60 | 29.33 | 140.10 | 1265.0 | 0.11780 | 0.27700 | 0.35140 | 0.15200 | 0.2397 | 0.07016 | … | 39.42 | 184.60 | 1821.0 | 0.16500 | 0.86810 | 0.9387 | 0.2650 | 0.4087 | 0.12400 | 0 |
| 568 | 7.76 | 24.54 | 47.92 | 181.0 | 0.05263 | 0.04362 | 0.00000 | 0.00000 | 0.1587 | 0.05884 | … | 30.37 | 59.16 | 268.6 | 0.08996 | 0.06444 | 0.0000 | 0.0000 | 0.2871 | 0.07039 | 1 |
5 rows × 31 columns
- In [4] :
df.describe()
- Out [4] :
| mean radius | mean texture | mean perimeter | mean area | mean smoothness | mean compactness | mean concavity | mean concave points | mean symmetry | mean fractal dimension | … | worst texture | worst perimeter | worst area | worst smoothness | worst compactness | worst concavity | worst concave points | worst symmetry | worst fractal dimension | class | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| count | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | … | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 | 569.000000 |
| mean | 14.127292 | 19.289649 | 91.969033 | 654.889104 | 0.096360 | 0.104341 | 0.088799 | 0.048919 | 0.181162 | 0.062798 | … | 25.677223 | 107.261213 | 880.583128 | 0.132369 | 0.254265 | 0.272188 | 0.114606 | 0.290076 | 0.083946 | 0.627417 |
| std | 3.524049 | 4.301036 | 24.298981 | 351.914129 | 0.014064 | 0.052813 | 0.079720 | 0.038803 | 0.027414 | 0.007060 | … | 6.146258 | 33.602542 | 569.356993 | 0.022832 | 0.157336 | 0.208624 | 0.065732 | 0.061867 | 0.018061 | 0.483918 |
| min | 6.981000 | 9.710000 | 43.790000 | 143.500000 | 0.052630 | 0.019380 | 0.000000 | 0.000000 | 0.106000 | 0.049960 | … | 12.020000 | 50.410000 | 185.200000 | 0.071170 | 0.027290 | 0.000000 | 0.000000 | 0.156500 | 0.055040 | 0.000000 |
| 25% | 11.700000 | 16.170000 | 75.170000 | 420.300000 | 0.086370 | 0.064920 | 0.029560 | 0.020310 | 0.161900 | 0.057700 | … | 21.080000 | 84.110000 | 515.300000 | 0.116600 | 0.147200 | 0.114500 | 0.064930 | 0.250400 | 0.071460 | 0.000000 |
| 50% | 13.370000 | 18.840000 | 86.240000 | 551.100000 | 0.095870 | 0.092630 | 0.061540 | 0.033500 | 0.179200 | 0.061540 | … | 25.410000 | 97.660000 | 686.500000 | 0.131300 | 0.211900 | 0.226700 | 0.099930 | 0.282200 | 0.080040 | 1.000000 |
| 75% | 15.780000 | 21.800000 | 104.100000 | 782.700000 | 0.105300 | 0.130400 | 0.130700 | 0.074000 | 0.195700 | 0.066120 | … | 29.720000 | 125.400000 | 1084.000000 | 0.146000 | 0.339100 | 0.382900 | 0.161400 | 0.317900 | 0.092080 | 1.000000 |
| max | 28.110000 | 39.280000 | 188.500000 | 2501.000000 | 0.163400 | 0.345400 | 0.426800 | 0.201200 | 0.304000 | 0.097440 | … | 49.540000 | 251.200000 | 4254.000000 | 0.222600 | 1.058000 | 1.252000 | 0.291000 | 0.663800 | 0.207500 | 1.000000 |
Convert to PyTorch Tensor
- In [5] :
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
- In [6] :
data = torch.from_numpy(df.values).float()
data.shape
- Out [6] :
torch.Size([569, 31])
- In [7] :
x = data[:, :-1]
y = data[:, -1:]
print(x.shape, y.shape)
torch.Size([569, 30]) torch.Size([569, 1])
- In [8] :
Train / Valid / Test ratio
ratios = [.6, .2, .2]
- In [9] :
train_cnt = int(data.size(0) * ratios[0])
valid_cnt = int(data.size(0) * ratios[1])
test_cnt = data.size(0) - train_cnt - valid_cnt
cnts = [train_cnt, valid_cnt, test_cnt]
print("Train %d / Valid %d / Test %d samples." % (train_cnt, valid_cnt, test_cnt))
Train 341 / Valid 113 / Test 115 samples.
- In [10] :
indices = torch.randperm(data.size(0))
x = torch.index_select(x, dim=0, index=indices)
y = torch.index_select(y, dim=0, index=indices)
x = x.split(cnts, dim=0)
y = y.split(cnts, dim=0)
for x_i, y_i in zip(x, y):
print(x_i.size(), y_i.size())
torch.Size([341, 30]) torch.Size([341, 1])
torch.Size([113, 30]) torch.Size([113, 1])
torch.Size([115, 30]) torch.Size([115, 1])
Preprocessing
- In [11] :
scaler = StandardScaler()
scaler.fit(x[0].numpy())
x =
[torch.from_numpy(scaler.transform(x[0].numpy())).float(), torch.from_numpy(scaler.transform(x[1].numpy())).float(),
torch.from_numpy(scaler.transform(x[2].numpy())).float()]
df = pd.DataFrame(x[0].numpy(), columns=cancer.feature_names)
df.tail()
- Out [11] :
| mean radius | mean texture | mean perimeter | mean area | mean smoothness | mean compactness | mean concavity | mean concave points | mean symmetry | mean fractal dimension | … | worst radius | worst texture | worst perimeter | worst area | worst smoothness | worst compactness | worst concavity | worst concave points | worst symmetry | worst fractal dimension | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 336 | 2.109400 | -0.982811 | 2.074373 | 2.141978 | 0.185909 | 0.834008 | 1.251482 | 1.990152 | 0.556192 | -0.173216 | … | 1.375889 | -0.629586 | 1.369703 | 1.215443 | -0.684062 | 0.122503 | 0.548355 | 1.215406 | -0.293150 | 0.080699 |
| 337 | 0.301591 | -0.605077 | 0.296273 | 0.197547 | -0.622822 | -0.197263 | -0.192340 | -0.226158 | -0.799453 | -0.451247 | … | 0.330800 | -0.888033 | 0.324745 | 0.218299 | 0.660655 | 0.418800 | 0.228829 | 0.124483 | 0.779932 | 0.641780 |
| 338 | -1.547870 | -0.808289 | -1.499389 | -1.213282 | 1.248526 | -0.454432 | -0.611624 | -0.789981 | -0.602723 | 1.873202 | … | -1.398695 | -0.988181 | -1.291901 | -1.070585 | 1.708544 | -0.164742 | -0.170464 | -0.184153 | 0.013892 | 0.866421 |
| 339 | -1.253992 | -0.846540 | -1.262700 | -1.041903 | -0.508002 | -1.105605 | -0.983191 | -0.925247 | 0.942498 | 0.008845 | … | -1.160364 | -0.784654 | -1.182241 | -0.941166 | 0.552717 | -1.025874 | -1.111026 | -1.000206 | -0.119264 | -0.175809 |
| 340 | -0.265338 | 0.317742 | -0.319287 | -0.319205 | -1.721811 | -0.996952 | -0.897690 | -0.964754 | -0.373802 | -0.589556 | … | -0.363802 | 0.439739 | -0.355996 | -0.405396 | -1.016868 | -0.598023 | -0.943469 | -1.147496 | -0.360512 | -0.565535 |
5 rows × 30 columns
Build Model & Optimizer
- In [12] :
# 입력 벡터 사이즈가 30차원
model = nn.Sequential(
nn.Linear(x[0].size(-1), 25), # 30~25차원
nn.LeakyReLU(),
nn.Linear(25, 20), # 25~20차원
nn.LeakyReLU(),
nn.Linear(20, 15), # 20~15차원
nn.LeakyReLU(),
nn.Linear(15, 10), # 15~10차원
nn.LeakyReLU(),
nn.Linear(10, 5), # 10~5차원
nn.LeakyReLU(),
nn.Linear(5, y[0].size(-1)),
nn.Sigmoid(),
)
model
- Out [12] :
Sequential(
(0): Linear(in_features=30, out_features=25, bias=True)
(1): LeakyReLU(negative_slope=0.01)
(2): Linear(in_features=25, out_features=20, bias=True)
(3): LeakyReLU(negative_slope=0.01)
(4): Linear(in_features=20, out_features=15, bias=True)
(5): LeakyReLU(negative_slope=0.01)
(6): Linear(in_features=15, out_features=10, bias=True)
(7): LeakyReLU(negative_slope=0.01)
(8): Linear(in_features=10, out_features=5, bias=True)
(9): LeakyReLU(negative_slope=0.01)
(10): Linear(in_features=5, out_features=1, bias=True)
(11): Sigmoid()
)
- In [13] :
optimizer = optim.Adam(model.parameters())
Train
- In [14] :
n_epochs = 10000
batch_size = 32
print_interval = 100
early_stop = 1000
- In [15] :
from copy import deepcopy
lowest_loss = np.inf
best_model = None
lowest_epoch = np.inf
- In [16] :
train_history, valid_history = [], []
for i in range(n_epochs):
indices = torch.randperm(x[0].size(0))
x_ = torch.index_select(x[0], dim=0, index=indices)
y_ = torch.index_select(y[0], dim=0, index=indices)
x_ = x_.split(batch_size, dim=0) # |xi| = (32,30)
y_ = y_.split(batch_size, dim=0) # |yi| = (32,1)
train_loss, valid_loss = 0, 0
y_hat = []
for x_i, y_i in zip(x_, y_):
y_hat_i = model(x_i)
loss = F.binary_cross_entropy(y_hat_i, y_i)
optimizer.zero_grad()
loss.backward()
optimizer.step()
train_loss += float(loss) # This is very important to prevent memory leak.
train_loss = train_loss / len(x_)
with torch.no_grad():
x_ = x[1].split(batch_size, dim=0)
y_ = y[1].split(batch_size, dim=0)
valid_loss = 0
for x_i, y_i in zip(x_, y_):
y_hat_i = model(x_i)
loss = F.binary_cross_entropy(y_hat_i, y_i)
valid_loss += float(loss)
y_hat += [y_hat_i]
valid_loss = valid_loss / len(x_)
train_history += [train_loss]
valid_history += [valid_loss]
if (i + 1) % print_interval == 0:
print('Epoch %d: train loss=%.4e valid_loss=%.4e lowest_loss=%.4e' % (
i + 1,
train_loss,
valid_loss,
lowest_loss,
))
if valid_loss <= lowest_loss:
lowest_loss = valid_loss
lowest_epoch = i
best_model = deepcopy(model.state_dict())
else:
if early_stop > 0 and lowest_epoch + early_stop < i + 1:
print("There is no improvement during last %d epochs." % early_stop)
break
print("The best validation loss from epoch %d: %.4e" % (lowest_epoch + 1, lowest_loss))
model.load_state_dict(best_model)
Epoch 100: train loss=5.6986e-03 valid_loss=2.1232e-01 lowest_loss=8.8456e-02
Epoch 200: train loss=1.7982e-05 valid_loss=3.7934e-01 lowest_loss=8.8456e-02
Epoch 300: train loss=4.0965e-06 valid_loss=4.2597e-01 lowest_loss=8.8456e-02
Epoch 400: train loss=1.6596e-06 valid_loss=1.1125e+00 lowest_loss=8.8456e-02
Epoch 500: train loss=7.0210e-07 valid_loss=1.1312e+00 lowest_loss=8.8456e-02
Epoch 600: train loss=3.9337e-07 valid_loss=1.1483e+00 lowest_loss=8.8456e-02
Epoch 700: train loss=1.7483e-07 valid_loss=1.1647e+00 lowest_loss=8.8456e-02
Epoch 800: train loss=9.1729e-08 valid_loss=1.1804e+00 lowest_loss=8.8456e-02
Epoch 900: train loss=4.5905e-08 valid_loss=1.1962e+00 lowest_loss=8.8456e-02
Epoch 1000: train loss=2.3287e-08 valid_loss=1.2109e+00 lowest_loss=8.8456e-02
There is no improvement during last 1000 epochs.
The best validation loss from epoch 12: 8.8456e-02
- Out [16] :
<All keys matched successfully>
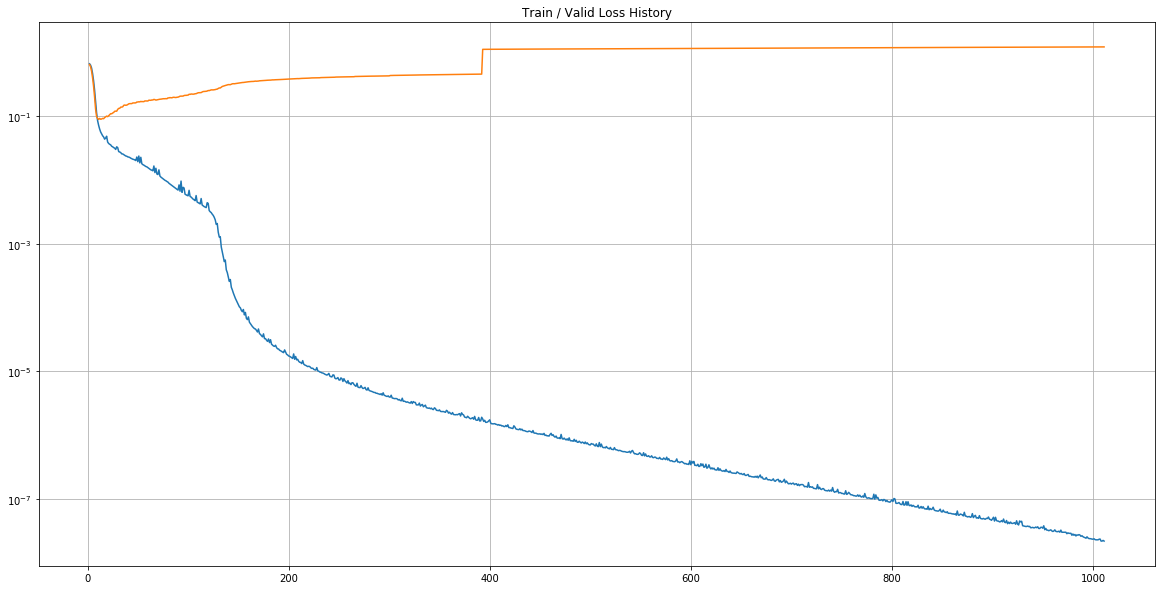
Loss History
- In [17] :
plot_from = 2
plt.figure(figsize=(20, 10))
plt.grid(True)
plt.title("Train / Valid Loss History")
plt.plot(
range(plot_from, len(train_history)), train_history[plot_from:],
range(plot_from, len(valid_history)), valid_history[plot_from:],
)
plt.yscale('log')
plt.show()
결과값 도출
- In [18] :
test_loss = 0
y_hat = []
with torch.no_grad():
x_ = x[2].split(batch_size, dim=0)
y_ = y[2].split(batch_size, dim=0)
for x_i, y_i in zip(x_, y_):
y_hat_i = model(x_i)
loss = F.binary_cross_entropy(y_hat_i, y_i)
test_loss += loss # Gradient is already detached.
y_hat += [y_hat_i]
test_loss = test_loss / len(x_)
y_hat = torch.cat(y_hat, dim=0)
print("Test loss: %.4e" % test_loss)
Test loss: 7.3890e-02
- In [19] :
correct_cnt = (y[2] == (y_hat > .5)).sum()
total_cnt = float(y[2].size(0))
print('Test Accuracy: %.4f' % (correct_cnt / total_cnt))
Test Accuracy: 0.9913
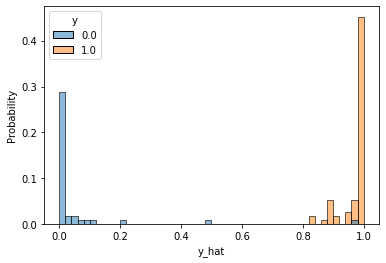
- In [20] :
df = pd.DataFrame(torch.cat([y[2], y_hat], dim=1).detach().numpy(),
columns=["y", "y_hat"])
sns.histplot(df, x='y_hat', hue='y', bins=50, stat='probability')
plt.show()
- In [21] :
from sklearn.metrics import roc_auc_score
roc_auc_score(df.values[:, 0], df.values[:, 1])
- Out [21] :
0.9951550387596899